Featured Publications
Work from Jon Brown Lab has recently been published in Nature, Circulation, Circulation Research, iScience, JCI Insight, JACC: Basic to Translational Science, Proceedings of the National Academy of Sciences.
Cardiomyocyte-restricted expression of progerin confers cardiac hypertrophy
Journal: Circulation Research
Year: 2025
PMID: 41064883
Hutchinson-Gilford progeria syndrome (HGPS) is a rapid, progressive aging disease caused by a mutation in the Lamin A gene (LMNA). Alternate splicing of LMNA produces a truncated mRNA and mutant protein called progerin, which exerts dominant negative effects and causes severe atherosclerosis.
Recent work in a human cohort has identified cardiac hypertrophy in children with HGPS, presumed to be a manifestation of systemic hypertension and vascular dysfunction. This study tested the hypothesis that progerin expression in cardiomyocytes can promote cardiac dysfunction. We expressed Progerin cDNA via cardiac troponin packaged in an AAV9 vector. Adult animals were injected at 8 weeks of age and followed for 9 months.
Progerin expression in cardiomycoytes promoted gradual, progressive cardiac hypertrophy, as measured by echocardiography with associated signs of diastolic dysfunction. Pathologic analysis revealed significant cardiomyocyte enlargement and increased DNA damage. These data demonstrate a cell automonomous effect of progerin expression in cardiomyocytes and uncover a novel role for this mutant protein in promoting cardiac aging.
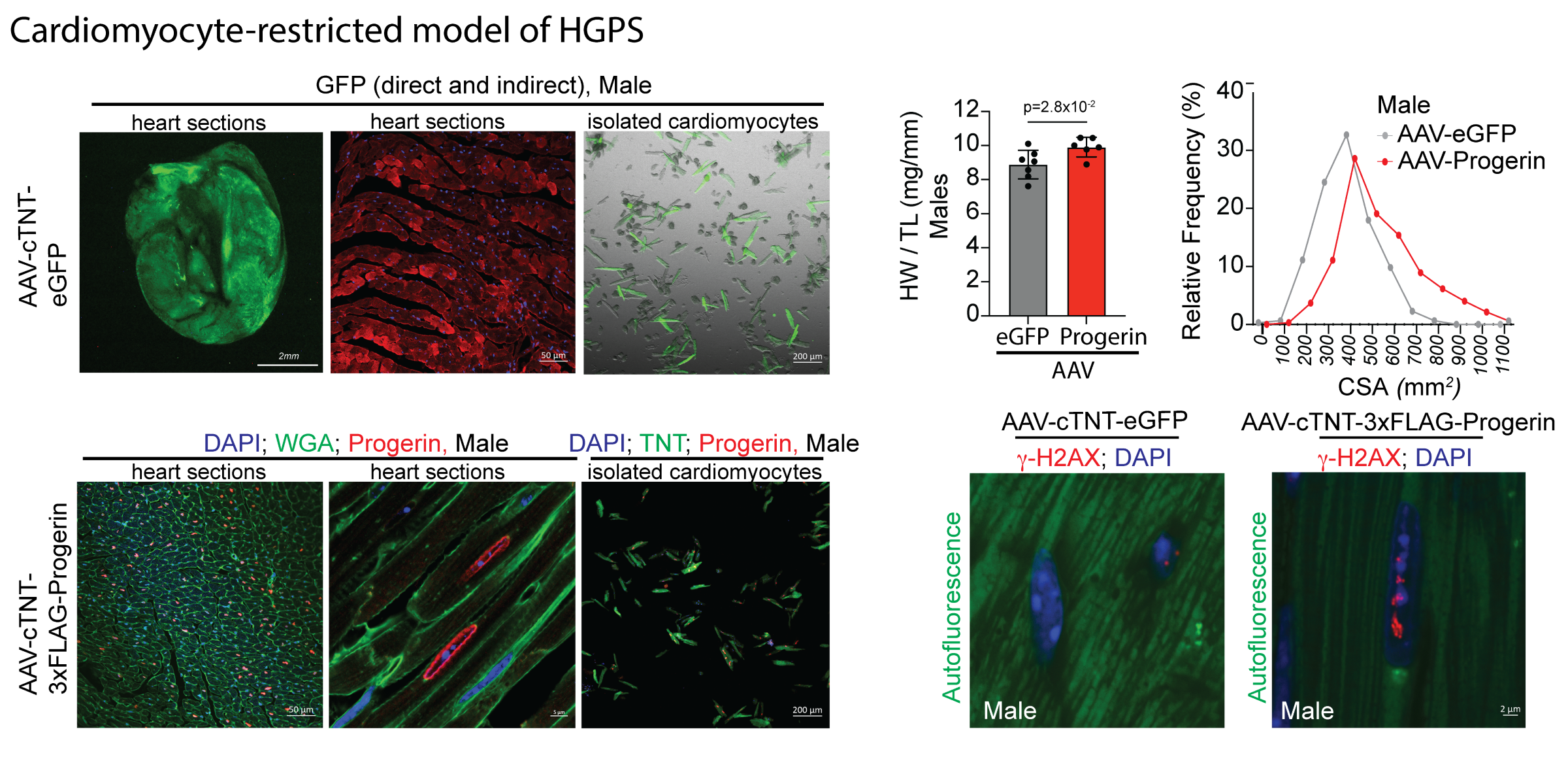
The images above depict a cardiomyocyte-restricted model of Hutchinson-Gilford progeria syndrome (HGPS).
Journal: Circulation
Year: 2025
PMID: 41052239
Peripheral artery disease (PAD) is a manifestation of atherosclerosis within the lower extremities that affects approximately 10 million people in the United States. Emerging evidence suggests that DNA methylation may play a critical role in the development and progression of cardiovascular disease. However, the relationship between DNA methylation and PAD remains poorly understood.
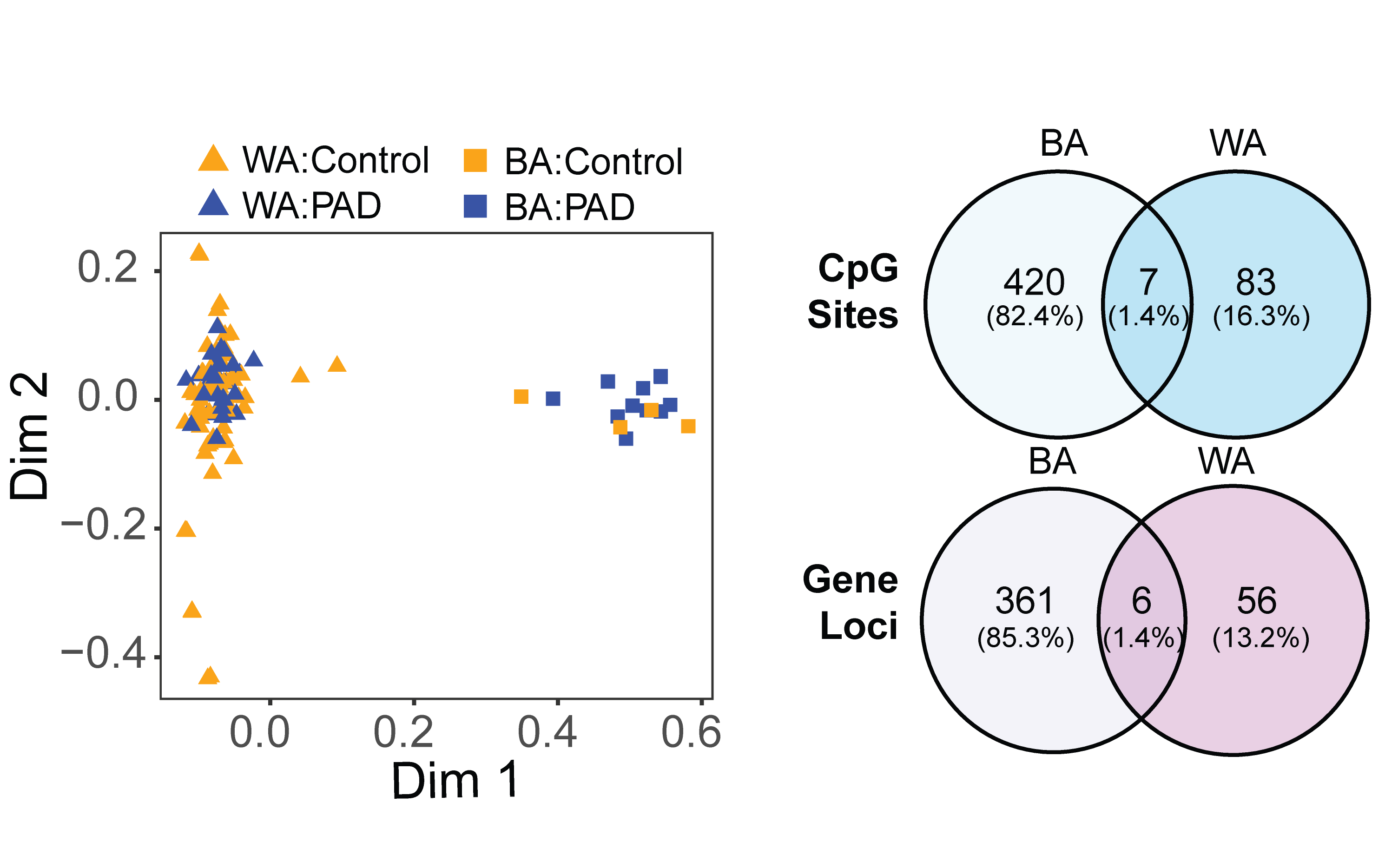
We conducted a cross-sectional analysis of baseline samples from a PAD intervention trial in which patients with intermittent claudication and an ankle-brachial index ≤ 0.85 and age-matched controls were recruited. Multidimensional Scaling (MDS) analysis of the 10,000 most variable CpG sites revealed clear separation by self-reported race.
These findings highlight the potential for novel determinants of PAD at the level of DNA methylation between these groups. Differences in PAD outcomes, particularly among Black Americans, are well-documented and have been attributed to health outcomes, as well as other environmental, behavioral, and clinical exposures that are more common among individuals who self-identify as Black. These exposures affect DNA methylation and represent a potential mediating pathway by which these stimuli affect disease risk.
The transcription factor ZNF469 regulates collagen production in liver fibrosis
Journal: JCI Insight
Year: 2025
PMID: 39998893
Metabolic dysfunction-associated steatotic liver disease (MASLD) - characterized by excess accumulation of fat in the liver - now affects one-third of the world's population. As MASLD progresses, extracellular matrix components including collagen accumulate in the liver, causing tissue fibrosis, a major determinant of disease severity and mortality.
To identify transcriptional regulators of fibrosis, we computationally inferred the activity of transcription factors (TFs) relevant to fibrosis by profiling the matched transcriptomes and epigenomes of 108 human liver biopsies from a deeply characterized cohort of patients spanning the full histopathologic spectrum of MASLD. CRISPR-based genetic KO of the top 100 TFs identified ZNF469 as a regulator of collagen expression in primary human hepatic stellate cells (HSCs). Gain- and loss-of-function studies established that ZNF469 regulates collagen genes and genes involved in matrix homeostasis through direct binding to gene bodies and regulatory elements.
By integrating multiomic large-scale profiling of human biopsies with extensive experimental validation, we demonstrate that ZNF469 is a transcriptional regulator of collagen in HSCs. Overall, these data nominate ZNF469 as a previously unrecognized determinant of MASLD-associated liver fibrosis.
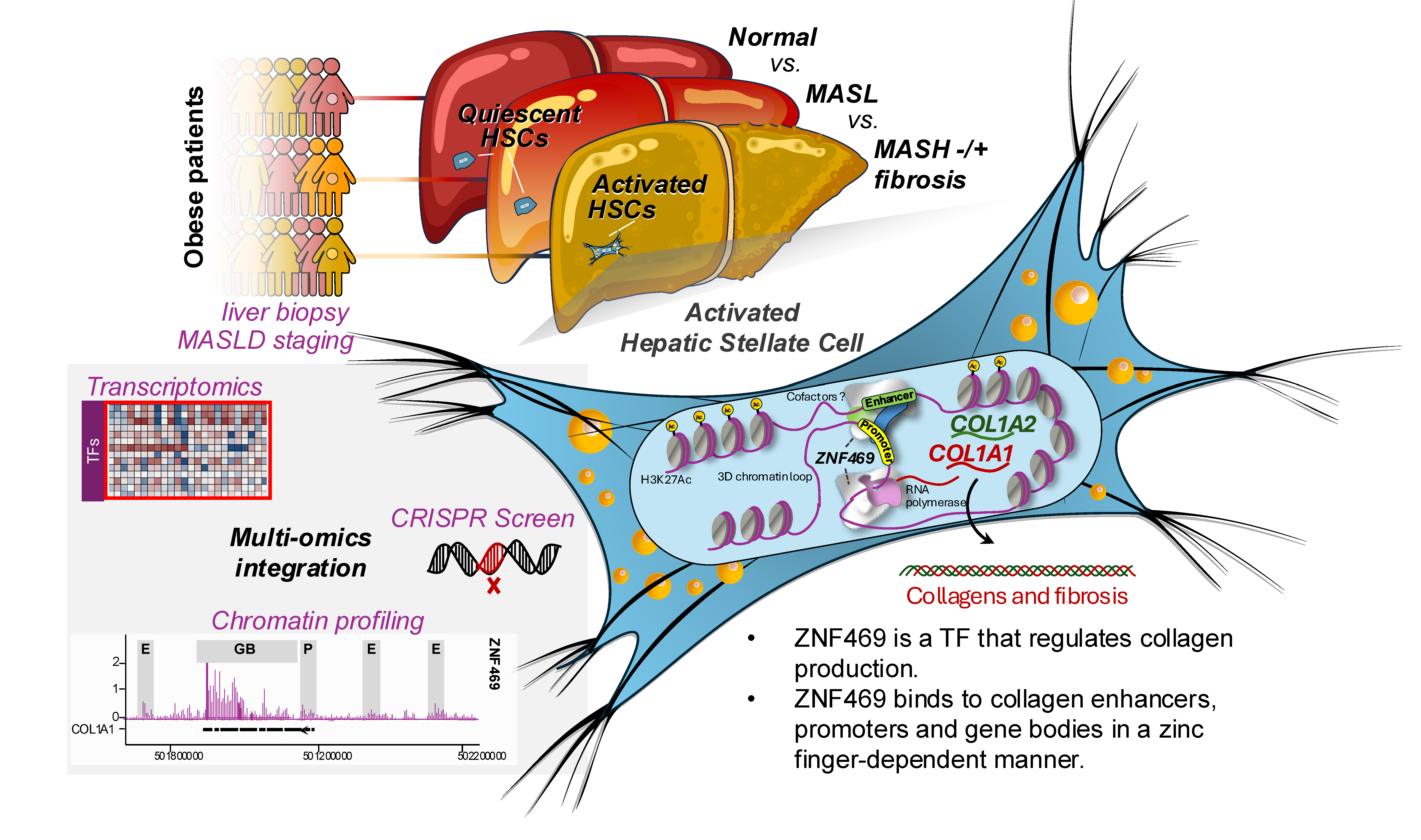
The graphical image above illustrates how the transcription factor ZNF469 regulates collagen production in liver fibrosis.
Journal: iScience
Year: 2024
PMID: 38868181
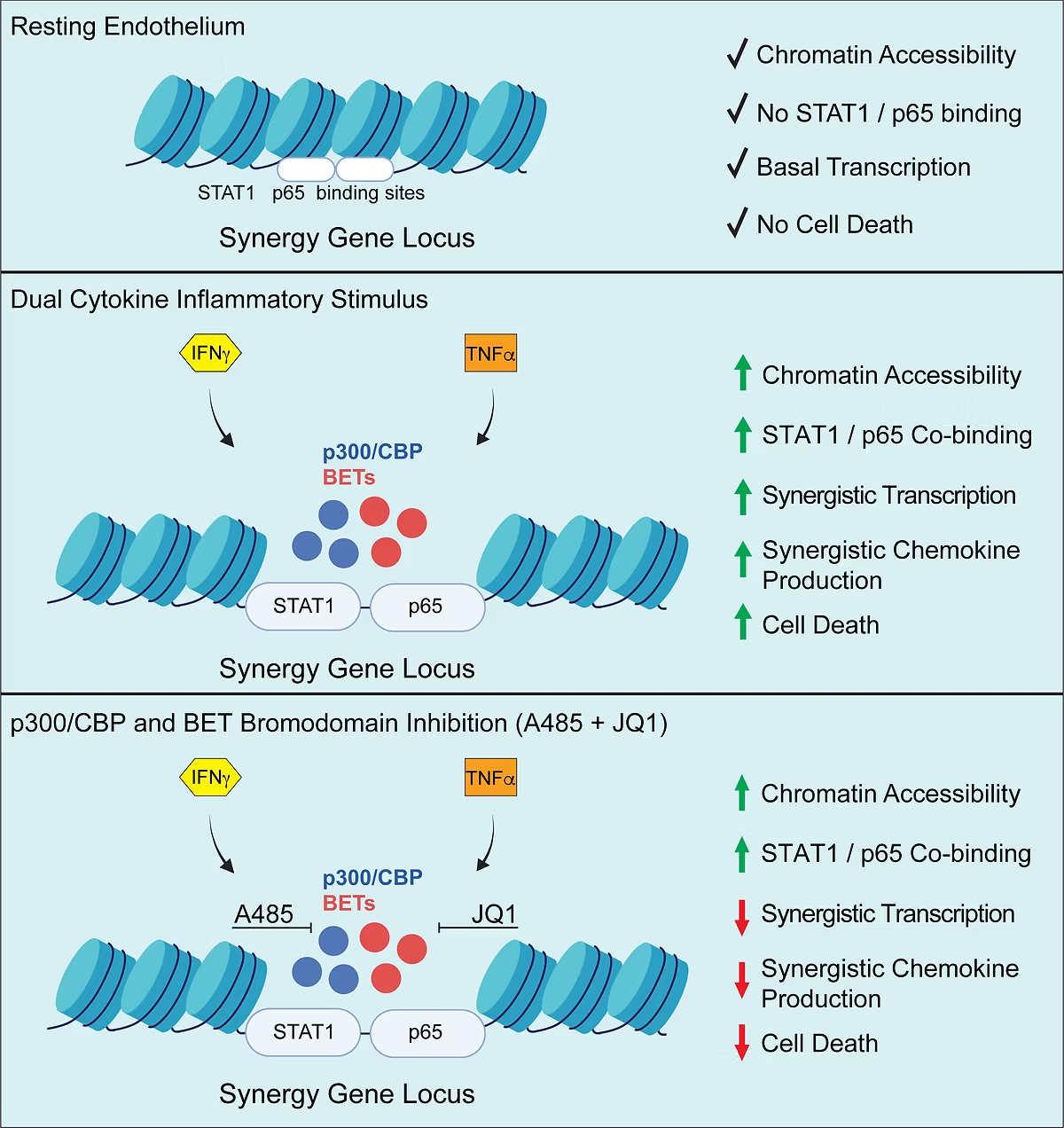
Combinatorial signaling by proinflammatory cytokines synergizes to exacerbate toxicity to cells and tissue injury during acute infections. To explore synergism at the gene regulatory level we investigated the dynamics of transcription and chromatin signaling in response to dual cytokines by integrating nascent RNA imaging mass spectrometry, RNA-sequencing, amplification-independent mRNA quantification, ATAC-sequencing and transcription factor profiling. Costimulation with interferon-gamma and tumor necrosis-alpha synergistically induced a small subset of genes, including the chemokines CXCL9, -10, -11. Gene induction coincided with increased chromatin accessibility at non-coding regions enriched for p65 and STAT1 binding sites.
To discover coactivator dependencies, we conducted a targeted chemogenomic screen of transcriptional inhibitors followed by modelling of inhibitor dose-response curves. These results identified high efficacy of either p300/CBP or BET bromodomain inhibitors to disrupt induction of synergy genes. Combination p300/CBP and BET bromodomain inhibition at subIC50 concentrations synergistically abrogated IFN/TNF-induced chemokine gene and protein levels.
Journal: Nature
Year: 2021
PMID: 33408413
Hutchinson-Gilford progeria syndrome (HGPS) is typically caused by a dominant-negative C•G-to-T•A mutation (c.1824 C>T, G608G) in LMNA, the nuclear lamin A gene. This mutation causes RNA mis-splicing that produces progerin, a toxic protein that induces rapid aging and shortens lifespan to ~14 years1-4. Adenine base editors (ABEs) perform targeted A•T-to-G•C base pair conversion with minimal byproducts and without requiring double-strand DNA breaks or donor DNA templates5,6.
Here, we describe the use of an ABE to directly correct the pathogenic HGPS mutation in cultured progeria patient-derived fibroblasts and in a mouse model of HGPS. Lentiviral delivery of ABE to patient-derived fibroblasts results in ~90% correction of the pathogenic allele, mitigation of RNA mis-splicing, reduced progerin levels, and correction of nuclear abnormalities. Unbiased off-target DNA and RNA analysis did not detect off-target editing activity in treated patient-derived fibroblasts. In transgenic mice homozygous for the human LMNA c.1824 C>T allele, a single retro-orbital injection of adeno-associated virus 9 (AAV9) encoding the ABE resulted in substantial, durable correction of the pathogenic mutation (~20-60% across various organs 6 months post-injection), restoration of normal RNA splicing, and reduction of progerin protein. In vivo base editing rescued vascular pathology, preserving vascular smooth muscle cell counts and preventing adventitial fibrosis. A single ABE AAV9 injection at P14 improved animal vitality and greatly extended median lifespan from 215 to 510 days.
These findings support the potential of in vivo base editing to treat HGPS, and other genetic diseases, by directly correcting the root cause of disease.
Journal: JACC: Basic to Translational Science
Year: 2019
PMID: 31768474
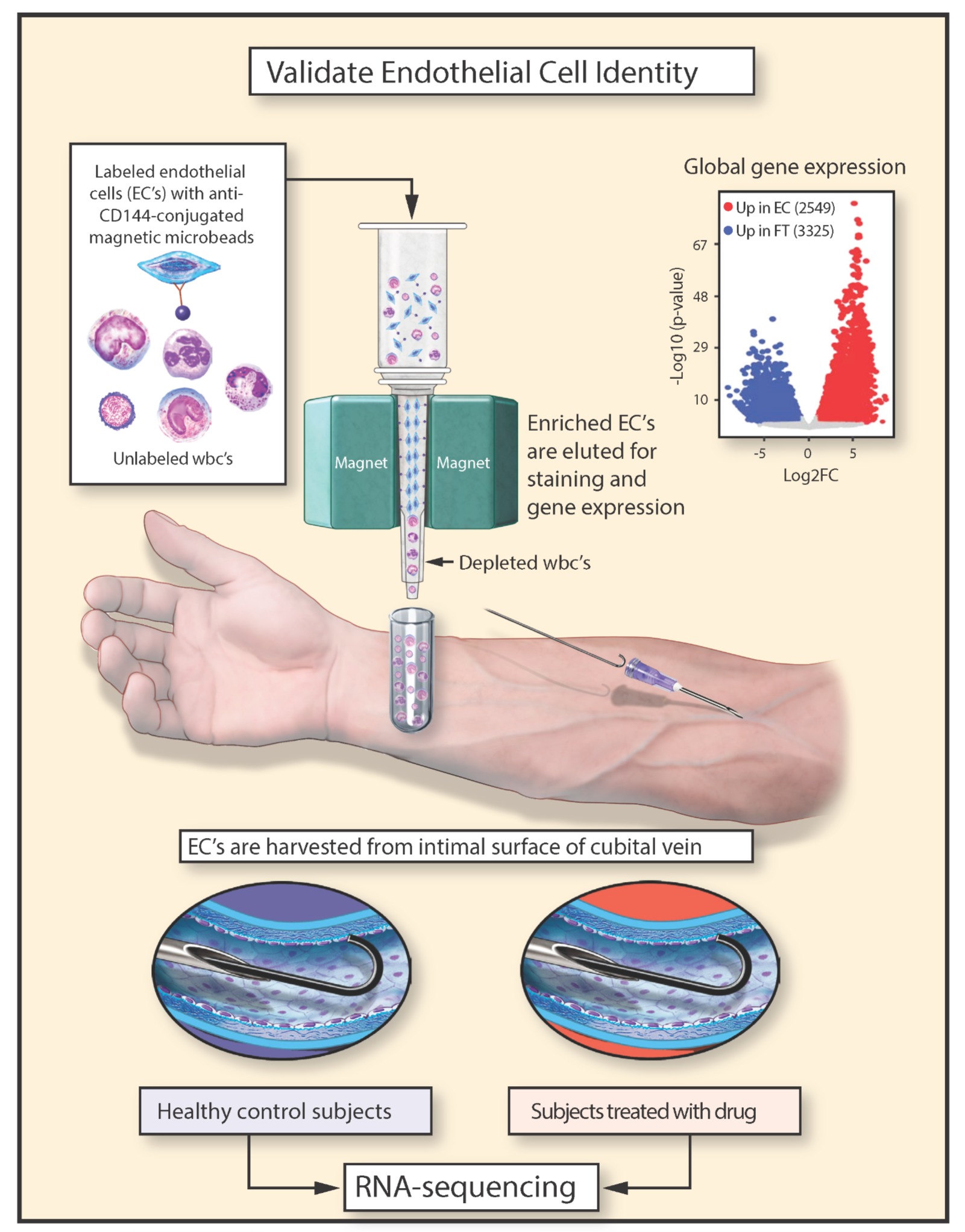
In this study we used low-input RNA-sequencing to annotate the molecular identity of human endothelial cells (ECs) isolated and immunopurified with CD144 microbeads.
After validating this technique, comparative gene expression profiling of ECs from healthy subjects and patients with type 2 diabetes mellitus identified both known and novel pathways linked with EC dysfunction. Modeling of diabetes by treating cultured ECs with high glucose identified shared changes in gene expression in diabetic cells.
Overall, the data demonstrate how purified ECs from patients can be used to generate new hypotheses about mechanisms of human vascular disease.
Journal: JCI Insight
Year: 2019
PMID: 31167964
Atherosclerotic plaques feature local proliferation of leukocytes and vascular smooth muscle cells (VSMCs) and changes in cellular metabolism.
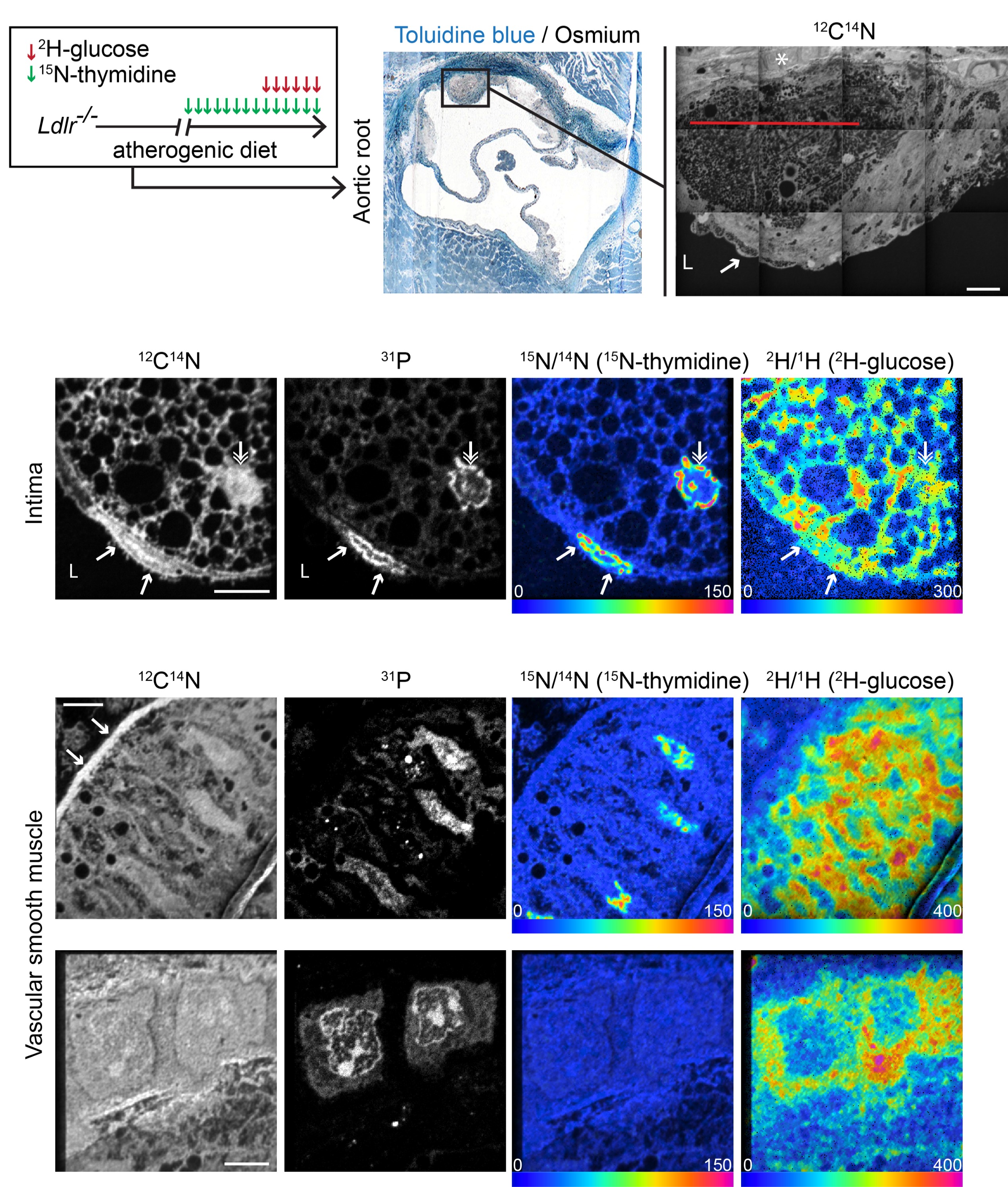
We used Multi-isotope Imaging Mass spectrometry (MIMS), a quantitative imaging platform, to measure coincident cell division and glucose utilization at suborganelle resolution in atherosclerotic plaques. In established plaques, 65% of intimal foam cells but only 4% of medial VSMCs were labeled with 15N-thymidine after 1 week of isotope treatment. Dividing cells demonstrated heightened glucose labeling. MIMS detected 2H-glucose label in multiple subcellular compartments within foam cells including lipid droplets, the cytosol and chromatin. Unexpectedly, we identified an intensely focal region of 2H-label in VSMCs underlying plaques. This signal diminished in regions of aorta without atherosclerosis. In advanced plaques, 15N-thymidine and 2H-glucose labeling in foam cells and VSMCs significantly decreased.
These data demonstrate marked heterogeneity in VSMC glucose metabolism that was dependent on both proliferative status and proximity of VSMCs to plaques. These results reveal how quantitative mass spectrometry coupled with isotope imaging can complement other methods used to study cell biology directly in the growing atherosclerotic plaque in vivo.